|
CLICK HERE TO READ ON IGB WEBSITE In the quest to decipher the evolutionary relationships of extinct organisms from fossils, researchers often face challenges in discerning key features from weathered fossils, or with prioritizing characteristics of organisms for the most accurate placement within a phylogenetic tree. Enter neural networks, sophisticated algorithms that underlie today’s image recognition technology. While previous attempts to utilize neural networks in classifying extinct organisms within phylogenetic trees have struggled, a new study, recently published in PNAS Nexus, heralds a significant breakthrough. The model has been trained to recognize and rank organism features based on known phylogenetic information, and can accurately place new organisms, including those that are extinct, within the intricate branches of evolutionary trees.
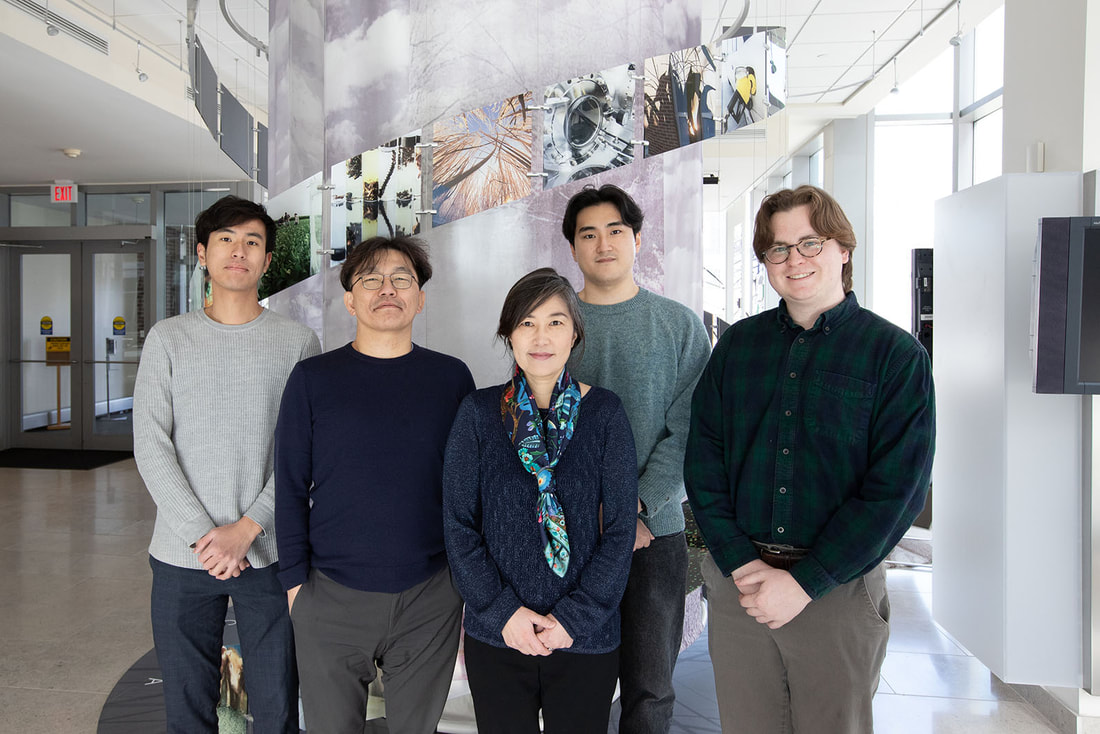
The team includes Surangi Punyasena (CAIM), an associate professor of Plant Biology at the University of Illinois Urbana-Champaign, Shu Kong, an assistant professor of science and technology at the University of Macau, and Marc-Élie Adaimé, a graduate student in Punyasena’s lab and first author on the study. According to Adaimé, the reason neural networks have trouble accurately classifying extinct organisms as opposed to living ones is often a matter of how they are trained. “Most paleontological AI studies typically focus on straightforward classification tasks, such as distinguishing between different fossil types,” explained Adaimé. “This approach works well within the scope of clearly defined categories, but less so with data that doesn’t fit these categories. Think of a model that has only been trained to classify images of dogs or cats. If it were presented with an image of a snake, the model would try to categorize it as either a dog or cat because it’s limited to what it was trained on. Similarly, there was no method previously that included phylogeny a priori into the model, so models did not learn to make sense of the features in an evolutionary or phylogenetic context. The goal of our research was to create a new modeling approach that would be trained on images in a phylogenetic context.” To accurately position organisms within a phylogenetic framework, neural networks must be trained not only to discern defining traits of various organism classes but also to recognize phylogenetic synapomorphies—derived features shared between organisms due to their common ancestry. This enables the network to determine the placement of organisms within a phylogenetic tree. The team chose to apply their model to the classification of pollen and spores —a ubiquitous and ancient entity found throughout the fossil record, with earliest fossils dating back hundreds of millions of years. The researchers first gathered optical superresolution images of modern and fossil pollen that had been taken at the Carl R. Woese Institute for Genomic Biology Core Facility. They trained their model using micrograph images of 30 extant (living) Podocarpus species. During this process, the model identified features it deemed important for classifying the pollen into different classes. Subsequently, these features were inputted into a secondary model, along with established phylogenetic data on the species, which then reweighted the features based on their phylogenetic significance. This approach enabled the model to generate a phylogenetically informed distance function, applicable to new pollen images provided to the model. To validate the model's efficacy, the researchers tested it on micrograph specimens of extinct pollen from Panama, Peru, and Columbia. While the exact phylogenetic relationships were not definitively known, paleoecologists had previously placed the pollen within Podocarpus based on morphological traits and geographical distribution. Impressively, the neural network model mirrored the placements made by the paleoecologists for nearly all specimens, underscoring its capacity to leverage morphological features learned during training to accurately position extinct species within a phylogenetic context. Punyasena noted that her lab is collaborating with colleagues at the Smithsonian National Museum of Natural History and the Smithsonian Tropical Research Institute to expand this work and apply it to a broader set of fossil pollen data. “International continental drilling projects are currently producing unimaginable amounts of fossil plant material,” said Punyasena. “Fully leveraging these new data sources means changing the way that we analyze and interpret fossil pollen. As a community, we need to take advantage of advances in deep learning and computer vision. This work demonstrates that the amount of evolutionary information captured in pollen morphology had been previously underestimated. The history of a plant species is captured in its shape and form. Machine learning allows us to discover these novel phylogenetic traits.” The researchers plan to enhance their model's accuracy and adaptability by expanding the sample size of images used for training. Furthermore, they aim to ensure the model remains current by integrating emerging advancements in machine learning. Adaimé emphasizes the model's versatility beyond pollen classification, foreseeing its potential application in categorizing various fossil organisms. “Machine learning models can make it easier to find features that are informative, because the way machine learning models think is obviously very different from what the way humans think,” said Adaimé. “It's going to be able to find patterns that make sense but probably aren’t intuitive to humans. And the benefit of this approach isn’t just limited to pollen, we expect these models will be generalizable to classifying fossils of other organisms as well.” The study was funded by the National Center for Supercomputing Applications and the University of Illinois, and can be found at https://doi.org/10.1093/pnasnexus/pgad419
0 Comments
In the realm of renewable energy where solar, wind, and thermal sources take center stage, a new technology is quietly emerging: energy generation through evaporation. Though in its infancy, evaporation technology holds the potential to revolutionize the clean energy sector by providing an efficient, abundant, and cost-effective alternative to fossil fuels.
Xi Chen, a professor with the ASRC’s Nanoscience Initiative, and his transdisciplinary research team have secured a $650,000 NSF Convergence Accelerator Phase 1 grant as part of the program’s Track M: Bio-Inspired Design Innovations. The funding will support the development of water-responsive materials and generators. The generators use the actuation created by these materials to harness evaporation and produce clean, renewable energy. “Our aim of this year-long, Phase 1 project is to establish platforms for the design and manufacturing of water-responsive materials and develop preliminary prototypes and simulation models of evaporation energy harvesting devices.” said Chen, the project’s principal investigator. The research team will produce materials from two sources. One is peptidoglycan, a macromolecule that can found within bacterial cell walls. Peptidoglycan strongly reacts to humidity, and it has the highest power and energy density compared to other actuator materials, including artificial muscles. The other material is silk collected from silkworm cocoons, which can be made into large sheets or other structures. As the water in these materials evaporate, the generators autonomously convert evaporation energy into mechanical motion and electricity. One notable advantage of this technology is its capacity to function both day and night, solving the important “intermittency” problem in energy storage for solar and wind. The ambitious research effort aims to produce prototypes within the next year and conduct product demonstrations for organizations and companies by the third year. Spanning various scientific disciplines within the ASRC and encompassing policy and marketing expertise, the transdisciplinary team also includes researchers from The City College of New York, Hunter College, Columbia University, New York University, National Renewable Energy Laboratory (NREL), GE research, Cannon, Ginkgo Bioworks, and ISEE Systems. The team’s diverse expertise is aimed at garnering public support for new approaches to renewable energy technology, said Charles Vörösmarty, founding director of the ASRC’s Environmental Science Initiative and a co-principal investigator on the project. “When you’re working with new technology, the general public isn’t going to know how it works and what it can offer,” said Vörösmarty. “For example, the Department Of Energy doesn’t have an evaporation program yet because the technology is so new. It’s important to gather people on our team with different expertise, like those in policy and marketing, so we can better inform people about the potential results the technology offers and bring it from the lab to real life to make it truly transformational.” The adaptable design of the generators allows them to be tailored to the needs and available space of their installation location, offering an alternative clean energy source for areas where solar and wind might face limitations due to weather or terrain. “One technique cannot solve all of the energy issues we’re facing today,” said Chen. “But this will provide another choice for clean energy that might be a better fit for people depending on their location and weather.” CLICK HERE TO READ ON IGB WEBSITE Cape lions used to roam the Cape Flats grassland plains of South Africa, in what is now known as Western Cape Providence. When Europeans arrived in South Africa in the mid-1600s, Cape lions, along with many other African carnivores and herbivores, were hunted as agricultural practice to protect livestock and humans. By the mid-1800s, less than 200 years since European arrival, Cape lions had been hunted to extinction. European naturalists described the Cape lion as having a particularly black mane and as being morphologically distinct. However, alternative depictions and descriptions of Cape lions from Indigenous people report mixed or light mane coloration. To shed light on this discrepancy, a recent study published in the Journal of Heredity, led by a team of researchers from the University of Illinois Urbana-Champaign, compared the genetic diversity and distinctiveness of Cape lions to modern lions across 13 African countries. The team features researchers from the Carl R. Woese Institute for Genomic Biology, including postdoctoral researcher and first-author Alida de Flamingh, Professor of Anthropology Ripan Malhi (CIS co-leader/GSP/GNDP/IGOH), Professor of Animal Sciences Alfred Roca (EIRH/GNDP), and Associate Professor of Integrative Biology Julian Catchen (CIS/GNDP), along with biologists from Roosevelt University and the Field Museum.
“What’s interesting is that the scientific name of the Cape lion, Panthera leo melanochaitus, literally means black mane, but this description was based on a single specimen,” said Julian Kerbis Peterhans, an emeritus professor at Roosevelt University. “Historically, we see lots of examples of creatures that are large and attractive like lions, where everybody wants to claim that they’d discovered a new one, without taking into account variation in the population, or whether that species is even unique.” Earlier investigations focused on limited segments of the Cape lion genome, offering the first indication that these lions might not be as distinct as initially believed. However, this study represents the first comprehensive examination of the entire Cape lion genome in comparison to contemporary lion populations across Africa. The team gathered samples from two Cape lion skulls currently housed at the Field Museum. These skulls were initially showcased at the South African Institute in Cape Town (1828-1838) as integral components of taxidermy mounts. The well-documented history of the skulls allowed the researchers to contextualize the diversity of the Cape lions within a specific time frame. “Unlike most other Cape Lion specimens around the world, these specimens had a traceable history and geographic collection location, so they had a pedigree,” said Thomas Gnoske, a Field Museum biologist. “As such, it was a great opportunity and challenge to see what application of the newest genomic methods could tell us about these specimens.” The genomic data collected from the skulls was compared to 118 existing mitogenomes and nuclear genomes of 53 other lions across Africa. Using complimentary genomic analyses, they found that the genome of the Cape lions was diverse and demonstrated genomic links with other lions from both the southern and eastern parts of Africa. While the researchers acknowledge the limitation of having only two Cape lion samples, they underscored that the results indicate that genetic characteristics that Cape lions had are still present in historic and some contemporary populations of lions in these regions in Africa. “One of the most surprising things was that we found such high genetic diversity in the Cape lion population,” said de Flamingh. “Both skulls were from the same small area, yet they had quite different mitochondrial DNA lineages and nuclear genomes. It tells us that there were very genetically diverse lions living in the Cape before they were extirpated.” The researchers also found that the Cape lion genomes exhibited high heterozygosity, and lacked traits commonly associated with small populations and inbreeding — characteristics frequently observed in endangered species facing population decline. This unexpected absence of such traits in the Cape lion genomes is particularly noteworthy, considering how close the skulls’ collection was to the species' extinction. “Contemporary species that are critically endangered and at high risk of extinction, such as the rhino or black-footed ferret, often have really small effective population sizes, which leads to inbreeding and lack of heterozygosity,” explained de Flamingh. “The Cape lions didn’t have any of those genomic signatures. What this means is that the Cape lions were hunted so rapidly that their genomes didn't have time to accumulate the signatures of long-term small population size.” The richness observed in the Cape lion genome implies that these lions likely exhibited significant phenotypic variation, including diverse mane coloration. This aligns more closely with alternative descriptions and Indigenous perspectives of the species, which the researchers say emphasizes the importance of incorporating diverse knowledge systems in enhancing our understanding of the histories of species. For me, the big takeaway from this study does not specifically have to do with Cape lions,” said Malhi. “The information we gained from the genomic data and analysis didn't fit what was thought about Cape lions based on colonial descriptions. This study is good example showing that identifying type specimens using information from people who are not originally from that area can result in ignoring diversity in a population that is important for understanding evolution.” The team suggests that this discovery provides insights for shaping future conservation strategies particularly for contemporary lion species in Africa. The results emphasize the importance of trans-country parks and heightened genetic connectivity between populations across Africa in order to maintain genetic diversity and flow. “Working with museums, such as the Field Museum, is an exciting opportunity to apply ancient DNA analyses to better understand human-animal interactions,” said de Flamingh. “I think it's an area that’s going to be studied more and more as our genetic technology continues to advance.” The study was funded by the USAID Wildlife TRAPS Project, USDA, NSF, and the University of Illinois. It can be found at https://doi.org/10.1093/jhered/esad081. CLICK HERE TO READ ON IGB WEBSITE Neurons intricately communicate and respond to stimuli within a vast network, orchestrating essential functions from basic bodily processes to complex thoughts. Traditional neuroscience methods, relying on in vivo electrophysiology (within a living organism), often have difficulty addressing the complexity of the brain as a whole. An alternative approach involves extracting cells from the organism and conducting studies on a culture dish instead (in vitro), providing researchers with enhanced control and precision in measuring neural processes. In a new study featured in Advanced Science, researchers unveil a cost-effective, open-source in vitro system for interfacing with neurons, offering a more accessible avenue for researchers interested in neural interactions. The study is part of a larger NSF funded project called Mind in Vitro, which explores how neurons interact with each other, not only to better understand the functions underlying complex systems like the brain, but also for the goal of eventually using in vitro neural networks for computation. To accomplish such a goal, the MiV project encompasses a highly interdisciplinary group of researchers, including those from computer science, engineering, neurobiology, physiology, and more.
“The goal of the MiV project is to ultimately use neurons for computation,” explained Zhi (Andrew) Dou, a graduate student in the Gazzola lab that helped lead the project. “This would allow for a system that is dynamic and constantly evolving, unlike traditional computing systems, and is actually more energy efficient as well.” The new MiV study, led by Mattia Gazzola (M-CELS), an associate professor of mechanical science and engineering, Xiaotian Zhang, a postdoctoral researcher in Gazzola’s lab, and Dou, describes an innovative approach to measuring neuron activity using micro-electrode array (MEA) technology. While there are commercial systems that utilize similar technology, they are often expensive and marketed towards specific experimental approaches. “The problem with the current technology for interfacing with neurons is that it’s mainly commercial systems, which are more standardized for specific testing conditions, generally within biochemistry or traditional neuroscience,” said Zhang. “Since we’re interested in engineering computing paradigms out of living cells, our motivation for this was to move away from the more standardized commercial systems and build something ourselves that we would have full control over.” In the new MiV apparatus, cells are placed onto a plate that contains MEAs, allowing the technology to interface with the neural substrates. The electrodes detect voltage from the neurons, and that voltage is amplified and sent to a computer, which then processes the data. Compared to the standard 60-electrode commercial system, the MiV system boasts over 500 electrodes, which allows for more data to be collected at once. The system also incorporates improved and/or novel features compared to commercial systems, such as portability, bi-directional communication with neurons, the ability to image neurons during recordings, and the ability to test multiple types of input (electric and optical) and multiple cell populations at a time. Despite these enhancements, the cost to build the MiV system is shockingly 10-times cheaper than the cost of a commercial system, and completely customizable. In order to further the dissemination of the system, the researchers have made their hardware and software models for the MiV system open source and freely available online. “Having our software and hardware be open source was really important to us,” said Zhang. “We wanted to bring the cost down to make this technology available to others, and have it be more flexible and versatile so that researchers can really customize their experimental settings for whatever they need. Our system is not for only one lab, or one group of people interested in one specific thing, it’s for everyone.” “We designed our system to be very easy to make and comprised of inexpensive parts, which will allow a lot of labs that can’t afford the commercial system to have their own system,” said Dou. “Even though the original design is for computational studies, we’ve made the structure easy to redesign and expand upon, so we’re pretty sure this will satisfy researchers no matter the kind of studies they want to do.” The researchers say other labs are already showing interest in the new technology for their own experiments. The MiV team plans to keep improving the design to make the hardware more automated and accessible, as well as explore ways to quantify and expand the technology’s performance. The study was funded by the NSF, and can be found at https://doi.org/10.1002/advs.202306826. Links for the open-source hardware designs and software can be found at https://gazzolalab.github.io/MiV-OH/ and https://miv-os.readthedocs.io/ respectively. CLICK TO READ ON CUNY WEBSITE
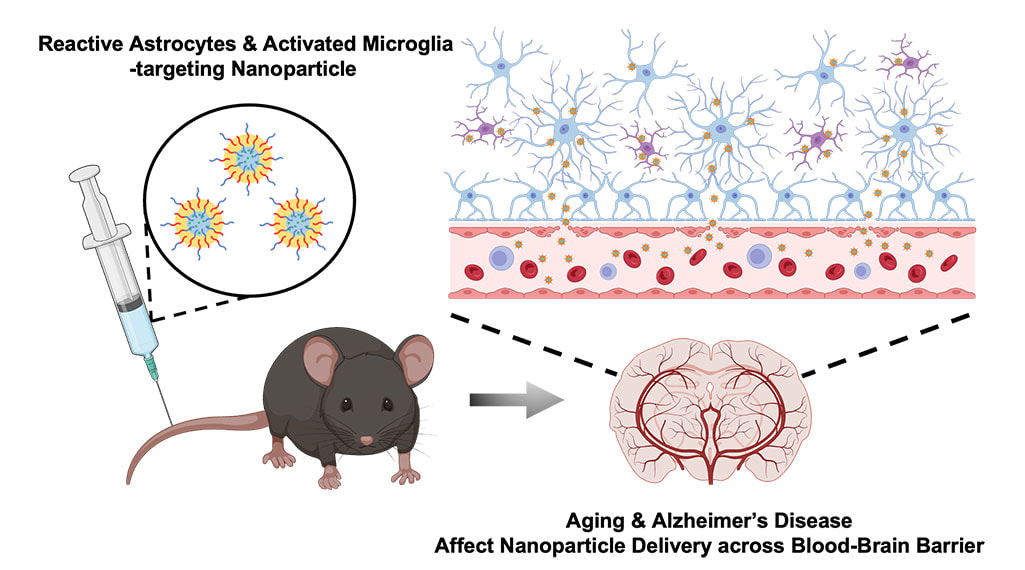
Traditional scientific methods for exploring the complexities of life break things down into smaller pieces and study how each part affects the big picture. Living systems are made up of billions of tiny interacting proteins, and scientists have made progress toward understanding the big picture by slowly putting together the pieces of the puzzle to comprehend each component. However, this process is reductionist, and according to Rein Ulijn, there’s unexplored potential in flipping this approach: assembling fundamental building blocks to forge a designed complex system from the ground up. To this end, Ulijn’s team aims to use mixtures of drastically simplified versions of proteins and observe how they exchange components and assemble to create properties like learning and memory. Ulijn, director of the Nanoscience Initiative at the Advanced Science Research Center at the CUNY Graduate Center and Einstein professor of chemistry at Hunter College, was recently awarded $675,000 from the Alfred P. Sloan Foundation to use this new approach to study whether synthetic biomolecular systems can be engineered to learn and acquire new traits or upgrade their performance in changing environments. To avoid getting lost in the complexity, Ulijn’s team will dramatically streamline the entire system by constructing simpler puzzles with much more basic versions of life’s molecules as the pieces. Their systems will be built from peptides, which are simplified versions of proteins that have specific interaction tendencies that lead to spontaneous self-organization. “Instead of studying one peptide at a time, we will mix and dynamically exchange them to form communities of interacting peptides,” Ulijn said. “By using a catalyst, we can enable peptides to exchange parts, like swapping words in a sentence. This allows the mixture to adapt and be amplified in response to changing conditions, which mimics learning. By also taking advantage of the aggregation of amplified peptides, sequence information can be stored and potentially retrieved later, which mimics memory.” The research project features three goals: creating a toolbox for designing peptides that, based on their specific interaction abilities, function as building blocks for adaptive biological systems; conducting experiments to demonstrate that systems can be trained and conditioned to selectively identify specific target compounds; and providing evidence that the system will be able to retain and recover information over time, reflecting a primitive form of learning and memory. To demonstrate memory, the researchers hope to be able to create designed peptide systems that distinguish between different flavor or fragrance molecules and show a stronger response when the same molecule is introduced a second or third time. “I am excited to be part of this newly funded initiative that explores the memory capabilities of peptides towards specific compounds,” said Kübra Kaygisiz, a newly recruited postdoctoral researcher in Ulijn’s lab. “This work will significantly enhance our understanding of the chemistry of life, and I look forward to advancing our knowledge in this fascinating field.” The project aims to improve understanding of how complex adaptive systems acquire new functions, and new findings could pave the way for innovations in information processing and memory in both biological and artificial systems. “Our approach allows us to study both the dynamically changing pieces and the system comprehensively,” Ulijn said. “We hope to achieve a deep understanding that empowers us to duplicate basic life-system features such as molecular adaptation, learning, and even memory in non-living systems.” CLICK HERE TO READ ON IGB WEBSITE Neurodegenerative disorders such as Alzheimer’s disease affect more than 270 million people worldwide. AD is the leading cause of dementia, resulting in memory loss due to atrophy of neurons in the hippocampus, which is the part of the brain that regulates learning and memory. Nanoparticles designed to carry drugs have emerged as a strategy for treating different diseases, but in the context of neurodegenerative disease, much of the research has focused on developing strategies for getting nanoparticles across the blood brain barrier and into targeted regions of the brain. In a new study, an interdisciplinary team of researchers at the University of Illinois Urbana-Champaign have developed nanoparticles that are able to selectively bind to activated astrocytes and microglia cells that mediate brain inflammation in AD, and found that both AD and aging strongly affect the ability of nanoparticles to cross the BBB and localize to the hippocampus. The BBB consists of a network of blood vessels surrounding the brain that tightly regulate what molecules (including drugs) can enter the brain. The BBB makes it difficult for nanoparticles carrying drugs to enter the brain, although nanoparticles can prevent the drugs from being “washed away”, or losing their activity along the way when passing through the BBB. However, research has suggested that the BBB weakens with AD and age. This inspired a team of researchers led by Joon Kong (M-CELS leader/EIRH/RBTE), a professor of chemical and biomolecular engineering, and Hee Jung Chung (M-CELS), an associate professor of molecular and integrative physiology, to synthesize a nanoparticle that could take advantage of this compromised BBB, and bind specifically to reactive astrocytes and microglia cells in the hippocampus of AD affected individuals. “I think people have overlooked how the vascular permeability of the BBB changes with AD pathology,” said Kong. “We thought, instead of putting peptides or proteins onto nanoparticles that can help them penetrate the BBB, as others have done, let’s just make the nanoparticles small enough that they can take advantage of the leaky BBB, and engineer these particles in a way that lets them remain in the brain in a stable manner.” The nanoparticles are designed to bind to CD44, a cell surface protein that is produced by reactive astrocytes and microglia cells, more than neurons, especially during neuroinflammation, a hallmark of AD-afflicted brain regions such as the hippocampus. The advantage of nanoparticles binding to these CD44-expressing cells is that the nanoparticles are retained longer in the hippocampus, rather than quickly being washed out, according to Kong. The researchers injected the CD44 seeking nanoparticles into both older and younger mice that either had AD or were healthy. They then looked at the distribution of nanoparticles in the hippocampus across the treatments. In the hippocampi of AD mice, they found high concentrations of nanoparticles regardless of age, though older AD mice had stronger concentrations than younger AD mice. The researchers say this was predicted, and further demonstrates that the BBBs of those with AD are considerably weakened. Not only were the nanoparticles able to penetrate the BBB, but they were also retained for longer in the hippocampus, for at least 2 hours post-injection, with preliminary data suggesting even longer retention. In the brains of healthy young mice, no nanoparticles were found, meaning that their BBBs were intact. However, to the team’s surprise, they found a significant amount of nanoparticles in the brains of healthy older mice, suggesting that the BBB weakens considerably with increasing age, even in those without AD. “This finding was surprising, because the older mice in this study are equivalent to a human age of only about 60 years old,” said Chung. “We knew there would be some leakiness of the BBB with age, but we thought there would be much less penetration of nanoparticles into the brain than we found. This highlights that there is age-dependent and disease-dependent penetration of the nanoparticles across the BBB to deep brain regions affected by AD.”
‘’This study offers valuable insights into advancing our understanding of nanoparticle transport to the brain in aging and Alzheimer's patients,” said Kai-Yu Huang, a graduate student in Kong’s lab. “It prompts us to think about the future strategies for the development of nano-scale drug carriers to target inflamed brain cells across different phases of aging-related brain disorders.’’ The next step is to try adding candidate drugs to the nanoparticles and see whether they could improve cognition and memory in AD mouse models, according to the researchers. They are also planning to measure how long their nanoparticles can be retained in the brain, which could help provide longer and more consistent drug delivery to patients treated with nanoparticles in the future. The team hopes this finding will provide a guideline for how to design drug carriers in the future for treating diseases, both within the brain and beyond. “This extends beyond just the brain, because this technology can be used for other diseases, not just brain disease,” said Chung. “By modifying the surface moiety of nanoparticles, we can directly target different organs, given we know something specific to target within those organs. The use of nanoparticles in medicine has broad and innovative applications.” This research was supported by the Alzheimer’s Disease Association, National Institutes of Health, the National Institute of Neurological Disorders and Stroke, the Chan Zuckerberg Biohub Chicago Acceleration Research Award, and the University of Illinois. The study can be found at https://doi.org/10.1021/acs.nanolett.3c03222 CLICK TO READ ON CUNY WEBSITE
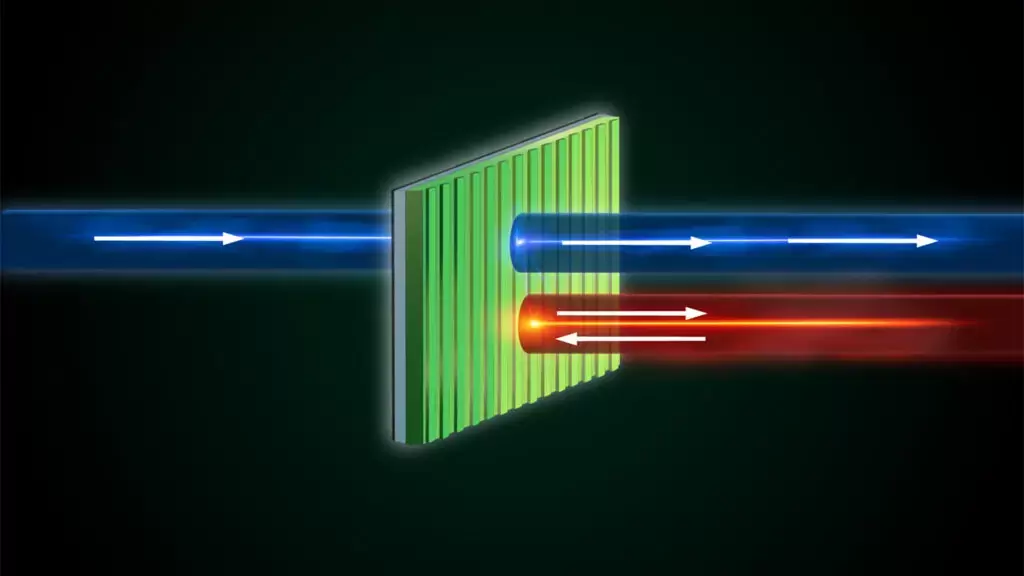
Using a thin layer of nano-patterned silicon, scientists are able to transmit light unidirectionally and prevent incoming reflections, all without the use of magnets. NEW YORK, December 8, 2023 — Much like the ebb and flow of ocean waves, light and sound waves bounce off objects and reflect back to their source. This inherent symmetry is known as reciprocity, and it can be problematic for the operation of many devices. For example, reflected waves from a powerful laser source can interfere with or even damage the device transmitting the signal. Special magnets, routers, and isolators, which are bulky and expensive, are normally needed to achieve asymmetrical wave propagation and overcome such problems. But a recently published study in Nature Photonics details a successful experiment conducted by a multi-institutional team of researchers that allowed them to break wave reciprocity without use of those devices. The work was led by Andrea Alù, founding director of the Photonics Initiative at the Advanced Science Research Center at the CUNY Graduate Center and distinguished professor of physics at the Graduate Center, and conducted in collaboration with colleagues at AMOLF in the Netherlands and Wayne State University. The team’s goal was to find a solution that could break light symmetry in strong laser beams propagating in free space without resorting to the use of magnetic materials, which often face equipment integration challenges. The approach focused on combining geometrical asymmetries with optical nonlinearities. These properties arise in materials that change their optical responses based on the intensity of the light flowing through them. Interestingly, the researchers found that the desired asymmetry could be achieved within a simple design and by using common materials. Their demonstration utilizes an ultrathin layer of silicon patterned at the nanoscale. “Silicon is the most common material you can think of in the context of optical technologies, and it is already the dominant material in so many of our devices and technologies,” said Alù. “We nanostructured a thin silicon film, only 100 nanometers in thickness or so, and demonstrated that, with the right shape and structure, we can largely enhance its optical non-linearity and leverage it to break reciprocity for light propagation.” The researchers started from a flat layer of silicon and etched lines onto it to achieve a sharp spectral response that helped enhance the otherwise weak nonlinearity of silicon. Crucially, the bottom part of the thin layer was left unetched in order to break the geometrical symmetry by just the right amount. Such geometrical asymmetry, combined with the enhanced nonlinearity, allows light to pass through when the film is hit from one side. Yet, when the patterned film is hit from the opposite side, the light is reflected away. In order to achieve this fit, the researchers engineered a “quasi-bound state in the continuum” in the device. Similar to an optical cavity made out of two mirrors, this state allows for light to be trapped in the thin engineered silicon film, enhancing its nonlinearity and intensity in a controllable fashion. “Our design is all about simplicity,” said Michele Cotrufo, assistant professor at the University of Rochester who was previously a postdoctoral researcher in Alù’s lab. “The idea of breaking reciprocity with tailored nonlinear materials has been floating around for some time, but no one was able to experimentally demonstrate it for light waves traveling in free space. We asked ourselves what the simplest, most practical design that still has all the right ingredients for the idea to work. Those ingredients for us were the patterning of a simple silicon film to create the required frequency dependence, vertical asymmetry to break mirror symmetry, and in-plane nanostructures to control the optical confinement. When we put these ingredients together, we realized that this was the best viable solution to demonstrate the working principle of this phenomenon.” While the current model operates within specific light bandwidths, the researchers aspire to create versatile models for different wavelengths, including those beyond optical frequencies. Exploring other nonlinear effects is also on the agenda, as the current material mostly relies on thermal phenomena to alter the refractive index. While these effects arise in a matter of microseconds, the team says it may not be fast enough for certain applications. Despite its limitations, the demonstrated silicon device holds vast potential applications. Beyond preventing reflections that are harmful to lasers, it may find uses in communication devices, such as cellphones and Lidar (the light equivalent of radar). Lidar is crucial in various technologies, including self-driving cars employing lasers to sense surrounding objects. The researchers aim to expand the reach of their silicon device, paving the way for widespread applications in diverse fields. The work was supported by the Air Force Office of Scientific Research, the Simons Foundation, and the Netherlands Organization for Scientific Research (NWO). CLICK HERE TO READ ON IGB WEBSITE A tiny but prolific world of microbes encompasses everything around us, both inside and out. Microbiomes, which are comprised of diverse communities of microbes, play a pivotal role in shaping human health, yet the intricacies of how different microbial compositions influence our well-being remains largely unknown. In a recent study published in PNAS, researchers at the University of Illinois Urbana-Champaign describe a new framework they have created to predict how species within microbiomes interact with each other to create unique compositions. “Microbes can be used in medicine, aka ‘bugs as drugs’, and these microbial therapeutics hold the possibility of being the answer to many of the diseases we face today,” said Shreya Arya, a graduate student in the O’Dwyer lab. “We’re trying to get away from using antibiotics to solve these issues because over time bacteria have developed antibiotic resistance. If the gut gets infected because of a pathogen, we want a way to be able to change the composition of the gut microbiome so that the gut microbiome is restored to a healthy state and the pathogen’s abundance is suppressed.”
Unfortunately, assessing interactions between each microbe species across diverse environments would demand an exponential volume of experimental data that would not be feasible to model. To overcome this hurdle, James O’Dwyer (CAIM), an associate professor of plant biology, along with Arya and Ashish George, a former postdoctoral researcher in O’Dwyer’s lab, sought to create a model that could predict outcomes of microbial communities based on the microbes initially present at the start. The model would provide a “landscape interaction value” between every microbe, that essentially characterizes how much effect each of the microbes has on another’s abundance and the microbiome’s outcome. “When doing this modeling, it’s important to ask the right question,” said O’Dwyer. “We could exhaustively try to model all of the pairwise comparisons and higher order interactions between species, which would give us the full dynamics of how the community changes over time, but that would literally take forever. Instead, we asked, at the end of all these microbial interactions, who's still there? How abundant are they and then what are their functions in that end-point community?” To address these questions, the research team started with a computational model capable of simulating microbial communities and predicting their outcomes. In this process, they uncovered a surprising revelation — most of the landscape interactions between microbes were near zero. This means that most microbes had a minimal impact on the final outcome of the microbiome, with only a select few playing a crucial role in predicting these outcomes. The researchers then used a method from the field of signal detection, called compressive sensing, which allows more information to be extracted from datasets with a sparse representation. The model was trained using existing microbiome datasets, and the results of these interactions were verified through real-world experiments with the same microbiomes to see if the resulting interactions and composition matched the predictions. Intriguingly, the researchers found that the “sparsity”, or abundance of zeros, of the landscape interactions held true, both in the model and in the real-world experiments. “I think there’s a lot we can learn about ecological communities in general from this,” O’Dwyer said. “We often think there's all these complex interactions, and this leads the structure and community functions to be hard to predict. But this shows that sometimes the outcomes are a bit simpler than you might expect. The magic here is that you don't have to learn everything about every initial condition through to every final state. You just have to learn a bit of it and it can give you enough information to know the whole thing.” The team is now interested in exploring why so many microbial landscape interactions were near zero, and trying larger datasets to see if that changes the patterns they found. “We want to understand why this sparsity is present in the first place, if that can tell us something about how microbiomes fundamentally are assembled, and how those species interact with each other,” explained Arya. “For example, even though a soil microbiome has very different species taxonomically speaking compared to the human gut, there may be similarities in the ways that microbial species interact with each other that we can predict based on the environment.” Arya hopes to continue fine-tuning the model so that it can be used to study particular microbiomes of interest, and accommodate more diverse datasets. One ultimate goal is to be able to use the model in personalized medicine, to help predict whether patients are at risk of certain pathogens establishing within their microbiomes compared to others.“In order to create microbial therapeutics, we need to understand which microbial species we need to combine in which environments in order to get the best function. And this is a first step towards that goal,” said Arya. This work was funded by the Simons Foundation, and can be found at https://doi.org/10.1073/pnas.2307313120 CLICK HERE TO READ ON CUNY WEBSITE
The new advance will enable pocket-sized devices that can perform detailed GPS-free precision navigation, medical imaging, food safety inspection and more. NEW YORK, NOVEMBER 9, 2023 — Lasers are essential tools for observing, detecting, and measuring things in the natural world that we can’t see with the naked eye. But the ability to perform these tasks is often restricted by the need to use expensive and large instruments. In a newly published cover-story paper in the journal Science, researcher Qiushi Guo demonstrates a novel approach for creating high-performance ultrafast lasers on nanophotonic chips. His work centers on miniaturizing mode-lock lasers — a unique laser that emits a train of ultrashort, coherent light pulses in femtosecond intervals, which is an astonishing quadrillionth of a second. Ultrafast mode-locked lasers are indispensable to unlocking the secrets of the fastest timescales in nature, such as the making or breaking of molecular bonds during chemical reactions, or light propagation in a turbulent medium. The high-speed, pulse-peak intensity and broad-spectrum coverage of mode-locked lasers have also enabled numerous photonics technologies, including optical atomic clocks, biological imaging, and computers that use light to calculate and process data. Unfortunately, state-of-the-art mode-locked lasers are currently expensive, power-demanding tabletop systems that are limited to laboratory use. “Our goal is to revolutionize the field of ultrafast photonics by transforming large lab-based systems into chip-sized ones that can be mass produced and field deployed,” said Guo, a faculty member with the CUNY Advance Science Research Center’s Photonics Initiative and a physics professor at the CUNY Graduate Center. “Not only do we want to make things smaller, but we also want to ensure that these ultrafast chip-sized lasers deliver satisfactory performances. For example, we need enough pulse-peak intensity, preferably over 1 Watt, to create meaningful chip-scale systems.” Realizing an effective mode-locked laser on a chip is not a straightforward process, however. Guo’s research leverages an emerging material platform known as thin-film lithium niobate (TFLN). This material enables very efficient shaping and precise control of laser pulses by applying an external radio frequency electrical signal. In their experiments, Guo’s team uniquely combined the high laser gain of III-V semiconductors and the efficient pulse shaping capability of TFLN nanoscale photonic waveguides to demonstrate a laser that can emit a high output peak power of 0.5 Watt. Beyond its compact size, the demonstrated mode-locked laser also exhibits many intriguing properties that are beyond reach by conventional ones, offering profound implications for future applications. For example, by adjusting the pump current of the laser, Guo was able to precisely tune the repetition frequencies of out pulses in a very wide range of 200 MHz. By employing the strong reconfigurability of the demonstrated laser, the research team hopes to enable chip-scale, frequency-stabilized comb sources, which are vital for precision sensing. Guo’s team will need to address additional challenges to realize scalable, integrated, ultrafast photonic systems that can be translated for use in portable and handheld devices, but his lab has overcome a major obstacle with this current demonstration. “This achievement paves the way for eventually using cell phones to diagnose eye diseases or analyzing food and environments for things like E. coli and dangerous viruses,” Guo said. “It could also enable futuristic chip-scale atomic clocks, which allows navigation when GPS is compromised or unavailable.” More information, including a copy of the paper, can be found online at the Science press package at https://www.eurekalert.org/press/scipak/ Bartering Light for Light: Scientists Discover New System to Control the Chaotic Behavior of Light11/2/2023 CLICK HERE TO READ ON CUNY WEBSITE
Studying how light beams interfere with each other in a stadium shaped arena gives researchers insight on its complex behavior. NEW YORK, November 2, 2023 -- Harnessing and controlling light is vital for the development of technology, including energy harvesting, computation, communications, and biomedical sensing. Yet, in real-world scenarios, complexity in light’s behavior poses challenges for its efficient control. Physicist Andrea Alù likens the behavior of light in chaotic systems to the initial break shot in a game of billiards. “In billiards, tiny variations in the way you launch the cue ball will lead to different patterns of the balls bouncing around the table,” said Alù, Einstein Professor of Physics at the CUNY Graduate Center, founding director of the Photonics Initiative at the CUNY Advanced Science Research Center and distinguished professor at CUNY. “Light rays operate in a similar way in a chaotic cavity. It becomes difficult to model to predict what will happen because you could run an experiment many times with similar settings, and you’ll get a different response every time.” In a new study published in Nature Physics, a team led by researchers at CUNY describe a new platform for controlling the chaotic behavior of light by tailoring its scattering patterns using light itself. The project was led by co-first authors Xuefeng Jiang, a former postdoctoral researcher in Alù’s lab who is now assistant professor of Physics with Seton Hall University, and Shixiong Yin, a graduate student in Alù’s lab. Conventional platforms for studying light’s behaviors typically use circular or regularly shaped resonant cavities in which light bounces and scatters in more predictable patterns. In a circular cavity, for example, only predictable and distinct frequencies (colors of light) survive, and each supported frequency is associated with a specific spatial pattern, or mode. One mode at a single frequency is sufficient to understand the physics at play in a circular cavity, but this approach does not unleash the full complexity of light behaviors seen in complex platforms, Jaing said. “In a cavity that supports chaotic patterns of light, any single frequency injected into the cavity can excite thousands of light patterns, which is conventionally thought to doom the chances of controlling the optical response,” Jaing said. “We have demonstrated that it is possible to control this chaotic behavior.” To address the challenge, the team designed a large stadium-shaped cavity with an open top and two channels on opposing sides that direct light into the cavity. As incoming light scatters off the walls and bounces around, a camera above records the amount of light escaping the stadium and its spatial patterns. The device features knobs on its sides to manage the light intensity at the two inputs, and the delay between them. Opposing channels cause the light beams to interfere with each other in the stadium cavity, enabling the control of one beam’s scattering by the other through a process known as coherent control—essentially, using light to control light, according to Alù. By adjusting the relative intensity and delay of the light beams entering the two channels, remarkably, researchers consistently altered the light’s radiation pattern outside the cavity. This control was enabled through a rare behavior of light in resonant cavities, called “reflectionless scattering modes” (RSMs), which had been theoretically predicted before but not observed in optical cavity systems. According to Yin, the ability to manipulate RSMs demonstrated in this work allows for the efficient excitation and control of complex optical systems, which has implications for energy storage, computing, and signal processing. “We found at certain frequencies our system can support two independen, overlapping RSMs, which cause all of the light to enter the stadium cavity without reflections back to our channel ports, thus enabling its control,” said Yin. “Our demonstration deals with optical signals within the bandwidth of optical fibers that we use in our daily life, so this finding paves a new way for better storage, routing, and control of light signals in complex optical platforms.” The researchers aim to incorporate additional knobs in future studies, offering more degrees of freedom to unravel further complexities in the behavior of light. “Coherent Control of Chaotic Optical Microcavity with Reflectionless Scattering Modes,” by Xuefeng Jiang, Shixiong Yin, Huanan Li, Jiamin Quan, Heedong Goh, Michele Cotrufo, Julius Kullig, Jan Wiersig, and Andrea Alù. |









 RSS Feed
RSS Feed
