|
CLICK HERE TO READ ON IGB WEBSITE Neurons intricately communicate and respond to stimuli within a vast network, orchestrating essential functions from basic bodily processes to complex thoughts. Traditional neuroscience methods, relying on in vivo electrophysiology (within a living organism), often have difficulty addressing the complexity of the brain as a whole. An alternative approach involves extracting cells from the organism and conducting studies on a culture dish instead (in vitro), providing researchers with enhanced control and precision in measuring neural processes. In a new study featured in Advanced Science, researchers unveil a cost-effective, open-source in vitro system for interfacing with neurons, offering a more accessible avenue for researchers interested in neural interactions. The study is part of a larger NSF funded project called Mind in Vitro, which explores how neurons interact with each other, not only to better understand the functions underlying complex systems like the brain, but also for the goal of eventually using in vitro neural networks for computation. To accomplish such a goal, the MiV project encompasses a highly interdisciplinary group of researchers, including those from computer science, engineering, neurobiology, physiology, and more.
“The goal of the MiV project is to ultimately use neurons for computation,” explained Zhi (Andrew) Dou, a graduate student in the Gazzola lab that helped lead the project. “This would allow for a system that is dynamic and constantly evolving, unlike traditional computing systems, and is actually more energy efficient as well.” The new MiV study, led by Mattia Gazzola (M-CELS), an associate professor of mechanical science and engineering, Xiaotian Zhang, a postdoctoral researcher in Gazzola’s lab, and Dou, describes an innovative approach to measuring neuron activity using micro-electrode array (MEA) technology. While there are commercial systems that utilize similar technology, they are often expensive and marketed towards specific experimental approaches. “The problem with the current technology for interfacing with neurons is that it’s mainly commercial systems, which are more standardized for specific testing conditions, generally within biochemistry or traditional neuroscience,” said Zhang. “Since we’re interested in engineering computing paradigms out of living cells, our motivation for this was to move away from the more standardized commercial systems and build something ourselves that we would have full control over.” In the new MiV apparatus, cells are placed onto a plate that contains MEAs, allowing the technology to interface with the neural substrates. The electrodes detect voltage from the neurons, and that voltage is amplified and sent to a computer, which then processes the data. Compared to the standard 60-electrode commercial system, the MiV system boasts over 500 electrodes, which allows for more data to be collected at once. The system also incorporates improved and/or novel features compared to commercial systems, such as portability, bi-directional communication with neurons, the ability to image neurons during recordings, and the ability to test multiple types of input (electric and optical) and multiple cell populations at a time. Despite these enhancements, the cost to build the MiV system is shockingly 10-times cheaper than the cost of a commercial system, and completely customizable. In order to further the dissemination of the system, the researchers have made their hardware and software models for the MiV system open source and freely available online. “Having our software and hardware be open source was really important to us,” said Zhang. “We wanted to bring the cost down to make this technology available to others, and have it be more flexible and versatile so that researchers can really customize their experimental settings for whatever they need. Our system is not for only one lab, or one group of people interested in one specific thing, it’s for everyone.” “We designed our system to be very easy to make and comprised of inexpensive parts, which will allow a lot of labs that can’t afford the commercial system to have their own system,” said Dou. “Even though the original design is for computational studies, we’ve made the structure easy to redesign and expand upon, so we’re pretty sure this will satisfy researchers no matter the kind of studies they want to do.” The researchers say other labs are already showing interest in the new technology for their own experiments. The MiV team plans to keep improving the design to make the hardware more automated and accessible, as well as explore ways to quantify and expand the technology’s performance. The study was funded by the NSF, and can be found at https://doi.org/10.1002/advs.202306826. Links for the open-source hardware designs and software can be found at https://gazzolalab.github.io/MiV-OH/ and https://miv-os.readthedocs.io/ respectively.
0 Comments
CLICK TO READ ON CUNY WEBSITE
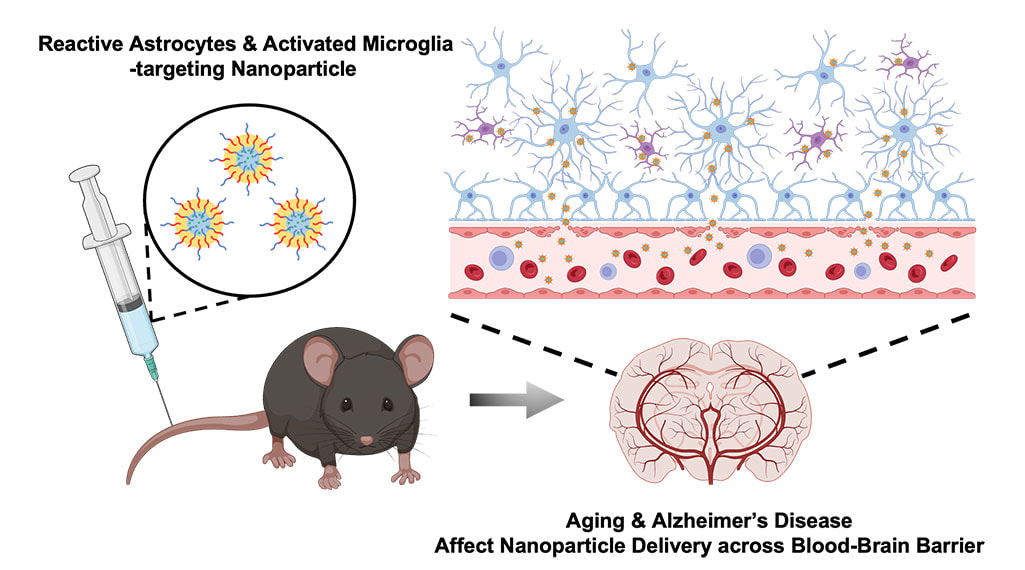
Traditional scientific methods for exploring the complexities of life break things down into smaller pieces and study how each part affects the big picture. Living systems are made up of billions of tiny interacting proteins, and scientists have made progress toward understanding the big picture by slowly putting together the pieces of the puzzle to comprehend each component. However, this process is reductionist, and according to Rein Ulijn, there’s unexplored potential in flipping this approach: assembling fundamental building blocks to forge a designed complex system from the ground up. To this end, Ulijn’s team aims to use mixtures of drastically simplified versions of proteins and observe how they exchange components and assemble to create properties like learning and memory. Ulijn, director of the Nanoscience Initiative at the Advanced Science Research Center at the CUNY Graduate Center and Einstein professor of chemistry at Hunter College, was recently awarded $675,000 from the Alfred P. Sloan Foundation to use this new approach to study whether synthetic biomolecular systems can be engineered to learn and acquire new traits or upgrade their performance in changing environments. To avoid getting lost in the complexity, Ulijn’s team will dramatically streamline the entire system by constructing simpler puzzles with much more basic versions of life’s molecules as the pieces. Their systems will be built from peptides, which are simplified versions of proteins that have specific interaction tendencies that lead to spontaneous self-organization. “Instead of studying one peptide at a time, we will mix and dynamically exchange them to form communities of interacting peptides,” Ulijn said. “By using a catalyst, we can enable peptides to exchange parts, like swapping words in a sentence. This allows the mixture to adapt and be amplified in response to changing conditions, which mimics learning. By also taking advantage of the aggregation of amplified peptides, sequence information can be stored and potentially retrieved later, which mimics memory.” The research project features three goals: creating a toolbox for designing peptides that, based on their specific interaction abilities, function as building blocks for adaptive biological systems; conducting experiments to demonstrate that systems can be trained and conditioned to selectively identify specific target compounds; and providing evidence that the system will be able to retain and recover information over time, reflecting a primitive form of learning and memory. To demonstrate memory, the researchers hope to be able to create designed peptide systems that distinguish between different flavor or fragrance molecules and show a stronger response when the same molecule is introduced a second or third time. “I am excited to be part of this newly funded initiative that explores the memory capabilities of peptides towards specific compounds,” said Kübra Kaygisiz, a newly recruited postdoctoral researcher in Ulijn’s lab. “This work will significantly enhance our understanding of the chemistry of life, and I look forward to advancing our knowledge in this fascinating field.” The project aims to improve understanding of how complex adaptive systems acquire new functions, and new findings could pave the way for innovations in information processing and memory in both biological and artificial systems. “Our approach allows us to study both the dynamically changing pieces and the system comprehensively,” Ulijn said. “We hope to achieve a deep understanding that empowers us to duplicate basic life-system features such as molecular adaptation, learning, and even memory in non-living systems.” CLICK HERE TO READ ON IGB WEBSITE Neurodegenerative disorders such as Alzheimer’s disease affect more than 270 million people worldwide. AD is the leading cause of dementia, resulting in memory loss due to atrophy of neurons in the hippocampus, which is the part of the brain that regulates learning and memory. Nanoparticles designed to carry drugs have emerged as a strategy for treating different diseases, but in the context of neurodegenerative disease, much of the research has focused on developing strategies for getting nanoparticles across the blood brain barrier and into targeted regions of the brain. In a new study, an interdisciplinary team of researchers at the University of Illinois Urbana-Champaign have developed nanoparticles that are able to selectively bind to activated astrocytes and microglia cells that mediate brain inflammation in AD, and found that both AD and aging strongly affect the ability of nanoparticles to cross the BBB and localize to the hippocampus. The BBB consists of a network of blood vessels surrounding the brain that tightly regulate what molecules (including drugs) can enter the brain. The BBB makes it difficult for nanoparticles carrying drugs to enter the brain, although nanoparticles can prevent the drugs from being “washed away”, or losing their activity along the way when passing through the BBB. However, research has suggested that the BBB weakens with AD and age. This inspired a team of researchers led by Joon Kong (M-CELS leader/EIRH/RBTE), a professor of chemical and biomolecular engineering, and Hee Jung Chung (M-CELS), an associate professor of molecular and integrative physiology, to synthesize a nanoparticle that could take advantage of this compromised BBB, and bind specifically to reactive astrocytes and microglia cells in the hippocampus of AD affected individuals. “I think people have overlooked how the vascular permeability of the BBB changes with AD pathology,” said Kong. “We thought, instead of putting peptides or proteins onto nanoparticles that can help them penetrate the BBB, as others have done, let’s just make the nanoparticles small enough that they can take advantage of the leaky BBB, and engineer these particles in a way that lets them remain in the brain in a stable manner.” The nanoparticles are designed to bind to CD44, a cell surface protein that is produced by reactive astrocytes and microglia cells, more than neurons, especially during neuroinflammation, a hallmark of AD-afflicted brain regions such as the hippocampus. The advantage of nanoparticles binding to these CD44-expressing cells is that the nanoparticles are retained longer in the hippocampus, rather than quickly being washed out, according to Kong. The researchers injected the CD44 seeking nanoparticles into both older and younger mice that either had AD or were healthy. They then looked at the distribution of nanoparticles in the hippocampus across the treatments. In the hippocampi of AD mice, they found high concentrations of nanoparticles regardless of age, though older AD mice had stronger concentrations than younger AD mice. The researchers say this was predicted, and further demonstrates that the BBBs of those with AD are considerably weakened. Not only were the nanoparticles able to penetrate the BBB, but they were also retained for longer in the hippocampus, for at least 2 hours post-injection, with preliminary data suggesting even longer retention. In the brains of healthy young mice, no nanoparticles were found, meaning that their BBBs were intact. However, to the team’s surprise, they found a significant amount of nanoparticles in the brains of healthy older mice, suggesting that the BBB weakens considerably with increasing age, even in those without AD. “This finding was surprising, because the older mice in this study are equivalent to a human age of only about 60 years old,” said Chung. “We knew there would be some leakiness of the BBB with age, but we thought there would be much less penetration of nanoparticles into the brain than we found. This highlights that there is age-dependent and disease-dependent penetration of the nanoparticles across the BBB to deep brain regions affected by AD.”
‘’This study offers valuable insights into advancing our understanding of nanoparticle transport to the brain in aging and Alzheimer's patients,” said Kai-Yu Huang, a graduate student in Kong’s lab. “It prompts us to think about the future strategies for the development of nano-scale drug carriers to target inflamed brain cells across different phases of aging-related brain disorders.’’ The next step is to try adding candidate drugs to the nanoparticles and see whether they could improve cognition and memory in AD mouse models, according to the researchers. They are also planning to measure how long their nanoparticles can be retained in the brain, which could help provide longer and more consistent drug delivery to patients treated with nanoparticles in the future. The team hopes this finding will provide a guideline for how to design drug carriers in the future for treating diseases, both within the brain and beyond. “This extends beyond just the brain, because this technology can be used for other diseases, not just brain disease,” said Chung. “By modifying the surface moiety of nanoparticles, we can directly target different organs, given we know something specific to target within those organs. The use of nanoparticles in medicine has broad and innovative applications.” This research was supported by the Alzheimer’s Disease Association, National Institutes of Health, the National Institute of Neurological Disorders and Stroke, the Chan Zuckerberg Biohub Chicago Acceleration Research Award, and the University of Illinois. The study can be found at https://doi.org/10.1021/acs.nanolett.3c03222 |



 RSS Feed
RSS Feed
